This AMOS plugin is made with the intent to simplify the creation of models based on pattern matrices in SPSS. It allows pasting the copied contents of a pattern matrix into a dialog box and constructs the applicable model in AMOS.

In another thread the topic of a Moodle Marketplace for the sale and purchase of themes and plugins is being not discussed. So I thought it might get discussed here. Right now there are almost no premium plugins for Moodle. Most plugin developers develop for themselves or their organisation, and quite often those are submitted to the Moodle plugins database.
Installation
Download
PatternMatrixBuilder.dll file from this repository's releases. When the download completes, copy the dll into AMOS' Plugins directory, which is normally located in C:Users{accountName}AppDataLocalAMOSDevelopmentAMOS{AMOSVersion}Plugins. The AppData folder is hidden by default, but you can access it by either typing in the path directly or showing hidden files.Restart AMOS, and the plugin should then show under the 'Plugins' menu in AMOS.
Usage
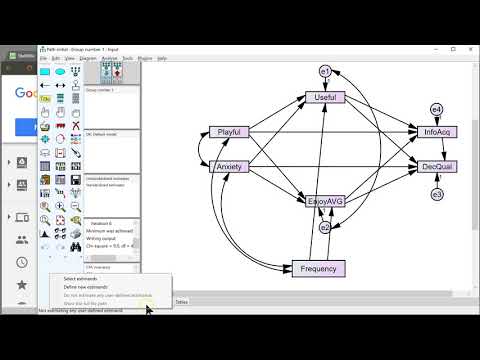
In AMOS, select the data set that corresponds to your pattern matrix. Click Plugins > Pattern Matrix Model Builder, select your pattern matrix in SPSS, Right click > Copy. In AMOS, paste your copied matrix into the input box. Then, click 'Create Diagram'. The model may take several minutes to generate, but could be quick depending on the size of your model.
Brucellosis is a zoonotic disease of high economic and public health importance worldwide [1, 2, 3]. It is caused by Brucella spp. and manifests itself as abortion and infertility in domestic and wild animal species and reduced milk production in cattle. In cattle the disease is mainly caused by B. abortus. However, other species of Brucella can also be isolated [4, 5, 6, 7, 8]. Brucellosis in humans is almost always associated with infected domestic and wild animals or their products and poses more risk to farmers, animal handlers, abattoir workers and veterinarians [9]. It causes a debilitating disease with unspecific symptoms comparable to other febrile conditions such as malaria, which may be chronically disabling. Treatment of human brucellosis is long and costly.
Brucella are small (0.5 to 0.7 by 0.6 to 1.5 μm), gram negative, non-motile, non- encapsulated, non-spore forming, rod shaped (coccobacilli) bacteria which are facultative intracellular parasites. The genus shows little variation genetically. To date there are 11 recognized species of Brucella which are genetically very similar although each has different host preferences [6]. Six are regarded as classical Brucella spp. Four members have recently been classified as additional species [10, 11] and recently the eleventh Brucella spp. has been described [12]. Three species are of great zoonotic and economic importance; these are B. abortus, B. melitensis, and B. suis which preferentially infect cattle, small ruminants and swine respectively. Some Brucella spp. are further divided into several biovars. So far, B. abortus has been subdivided into biovars 1, 2, 3, 4, 5, 6, 7 and 9 [13]. Several biovars of B. melitensis (biovar 1, 2, 3) and B. suis (biovar 1, 2, 3, 4, 5) are also recognized [14]. Brucella abortus biovar 1 accounts for more than 80 % of the total number of isolates worldwide whereas in Africa B. abortus biovar 3 has been reported in most of the few published studies [2, 4].
Screening of brucellosis can be performed by serological methods detecting antibodies directed against epitopes associated with the smooth lipopolysaccharide (S-LPS) [5]. Confirmation of the infection is done by culture and isolation of the bacteria. However, this bacterium is difficult to grow and the procedure is time consuming. Furthermore, the procedure poses a risk to laboratory personnel and should be performed in biosafety level 3 laboratories. Nevertheless this method remains the “Gold standard” for diagnosis of brucellosis and Brucella infections. Biotyping of Brucella spp. provides additional information. Polymerase Chain Reaction (PCR) and other molecular techniques have been developed and have found diagnostic application [1]. Detection of Brucella spp. or its DNA provide the only certain diagnosis [5]. Genotyping of Brucella spp. can be achieved by Multiple Loci Variable Number of Tandem Repeats Analysis (MLVA-VNT) which shows a very good discriminatory power [14]. Such data can provide molecular epidemiological information for elucidating transmission pattern.
Brucellosis is widely spread in African countries [2, 3, 15, 16]. Serological studies done in different parts of Tanzania indicate that the infection is widely spread in domestic animals, wildlife and human beings [17, 18]. In Tanzania the problem is bigger in pastoral systems and wildlife than in the dairy farming system [18]. Data on isolation of Brucella spp. both in humans and animals, with further characterization is scarce. Isolation of B. abortus and B. melitensis from cattle and small ruminants in Tanzania was reported more than 50 years ago. However characterization of the isolates was not performed [19]. Brucella melitensis and B. abortus have been isolated and characterized from cattle in Uganda and Kenya. In Tanzania, similar studies need to be performed to trace back the reservoir host species [7, 20]. It is not known whether cattle in Tanzania are infected with B. abortus or B. melitensis or both. Successful control of brucellosis requires knowledge of its epidemiology in different animal species and the circulating strains in the region have to be assessed. This information is scarce in Tanzania.
Therefore the aims of the present study were to investigate a dairy herd experiencing abortion in order to:- Establish within-herd prevalence of Brucella seropositive animals.
- Isolate, identify and characterize Brucella spp. from milk and abortion materials.
- Compare molecular characteristics of the obtained isolates with other strains in the region and outside.
- Investigate a possible spillover to small ruminants and dogs as they are a potential source of infection to cattle and can as well acquire infection from cattle.
